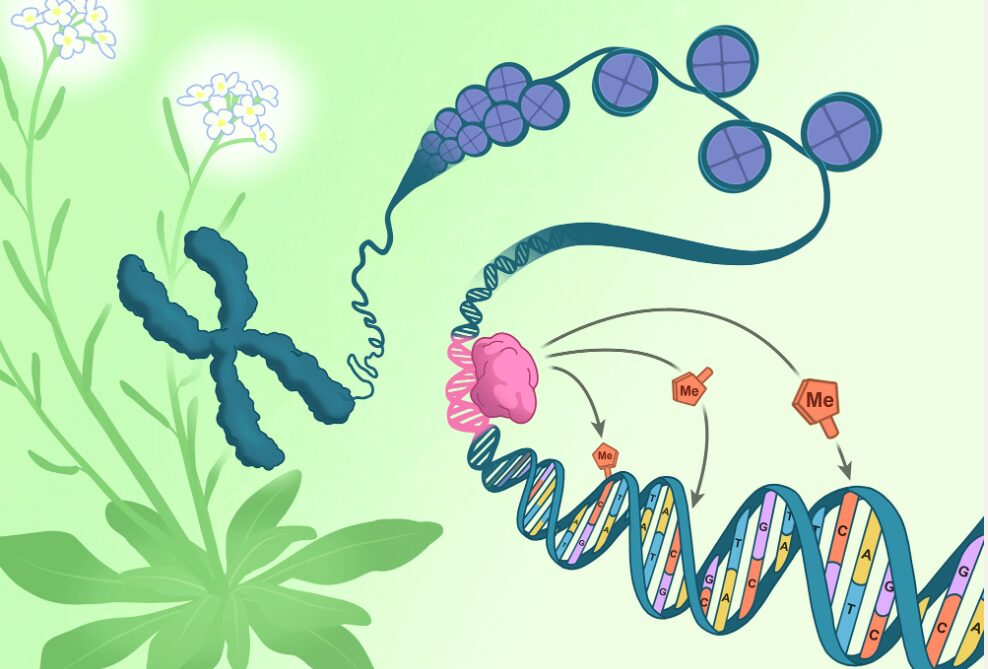
A chromosome pulled from the flowers of Arabidopsis thaliana (green and white) unspools to reveal DNA (blue) coiled around packaging-proteins called histones (purple). The direction of epigenetic changes by genetic features begins as the RIM transcription factor (pink) docks on a corresponding DNA sequence (pink). Once docked, the RIM transcription factor directs methylation machinery to tack methyl groups (orange) onto specific nearby cytosines (orange).
Click here for a high-resolution image.
Credit: Salk Institute
All the cells in an organism have the exact same genetic sequence. What differs across cell types is their epigenetics—meticulously placed chemical tags that influence which genes are expressed in each cell...
Read More






Recent Comments