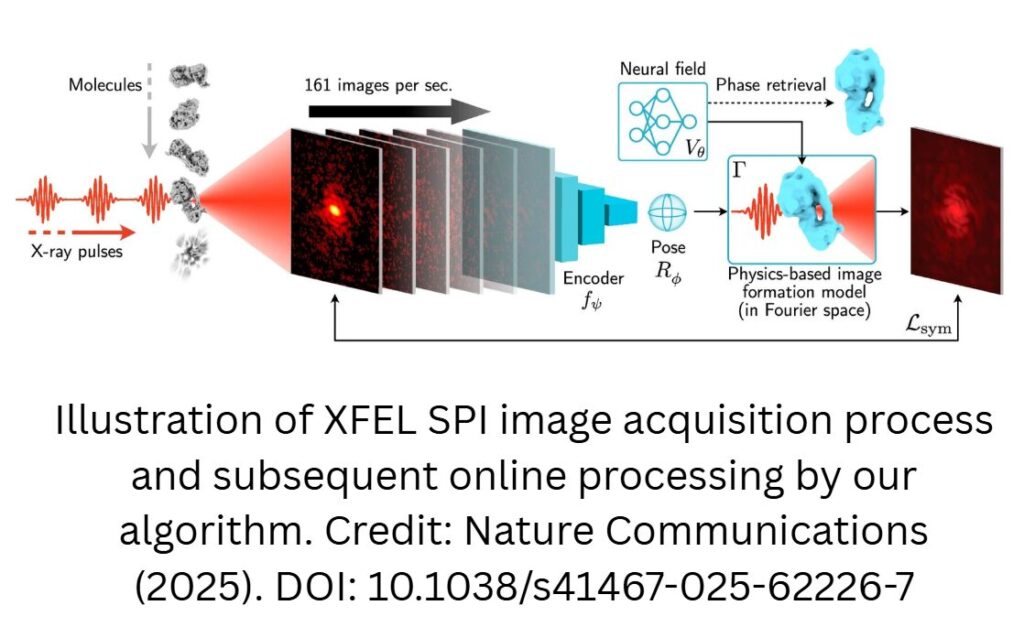
Soon, researchers may be able to create movies of their favorite protein or virus better and faster than ever before. Researchers at the Department of Energy’s SLAC National Accelerator Laboratory have pioneered a new machine learning method—called X-RAI (X-Ray single particle imaging with Amortized Inference)—that can “look” at millions of X-ray laser-generated images and create a three-dimensional reconstruction of the target particle. The team recently reported their findings in Nature Communications.
X-RAI’s ability to sort through a massive number of images and learn as it goes could unlock limits in data-gathering, allowing researchers to see molecules up close—and perhaps even on the move. “There is really no limit” to the dataset size it can handle, said SLAC staff scientist Frédéric Poitevin, one of the study’s principal investigators.
Lagging algorithms
The Linac Coherent Light Source (LCLS) at SLAC takes X-ray snapshots of atoms and molecules at work, revealing fundamental processes in materials, technology and living things.
The work took place at SLAC’s Linac Coherent Light Source (LCLS)—the world’s most powerful X-ray free-electron laser, where researchers blast samples with X-ray pulses just a few millionths of a billionth of a second long to gain insights into the structures and motions of molecules, such as proteins or viruses.
When researchers send these pulses through a sample, the X-rays scatter off the molecules in that sample and create a 2D scattering image on a detector. By combining many of those 2D images, capturing information about the molecules at many different angles, the researchers can reconstruct the molecules’ 3D structure.
But this process is a time-consuming task, requiring hundreds of thousands to millions of 2D images. On top of that, the algorithms traditionally used to piece scattering images together get slower as X-ray laser data pile up. For each snapshot, the algorithms try to predict the object’s 3D structure—the more snapshots, the more time spent puzzling over the reconstruction.
Such computing delays can limit researchers at LCLS, who may only have a few days of access to the facility to run a whole suite of experiments. “How could we make this faster to the point that we could actually do the reconstruction as we collect the data and not wait hours or days for the results?” asked Poitevin.
Unlocking LCLS datasets
To solve this problem, Poitevin and colleagues, including Gordon Wetzstein’s lab at Stanford University, developed a new machine learning program that processes X-ray laser data on the fly—and improves as it proceeds.
The neural network “looks” at the 2D images and predicts a 3D orientation of the sample particle. It also works in the opposite direction, taking a 3D projection and generating 2D images from it. This bidirectional process allows for the AI to continually refine the relationship between the 2D X-ray laser data and the 3D reconstruction. The more data, the better it understands the relationship between the 2D images and 3D structure, becoming more efficient.
For large datasets, X-RAI is “much faster” than other programs, said Jay Shenoy, the study’s first author and a Stanford Ph.D. student in computer science. In the paper, the team demonstrated the new algorithm could process up to 160 images per second in real time while predicting the object’s 3D structure on the computer screen.
The researchers also compared how well X-RAI reconstructed the 3D structure of two biomolecules—a ribosomal subunit and the protein ATP synthase—compared to two other algorithms and found that the new program produced reconstructions that were sharper.
The team hopes the advance will enable users to make the most of their time at LCLS and other X-ray lasers around the world. Experimental time at this cutting-edge X-ray light source is highly competitive, and successful applicants are only granted a few days of access at a time. X-RAI could help researchers get more out of the allocated time.
Opening the door to a speedy analysis of near-limitless amounts of data may also help researchers to study particles in motion. With enough images, they could reconstruct a movie of, for example, an enzyme interacting with a drug. With the improvements to X-ray imaging, “you might be able to actually get a more accurate picture of how your molecule moves,” said Poitevin. https://techxplore.com/news/2025-11-machine-algorithm-rapidly-reconstructs-3d.html






Recent Comments